Sex biases in ILC1 phenotypes do not alter immune control of cytomegalovirus
RM: https://orcid.org/0000-0002-1549-9663
CB: https://orcid.org/0000-0001-7816-656X
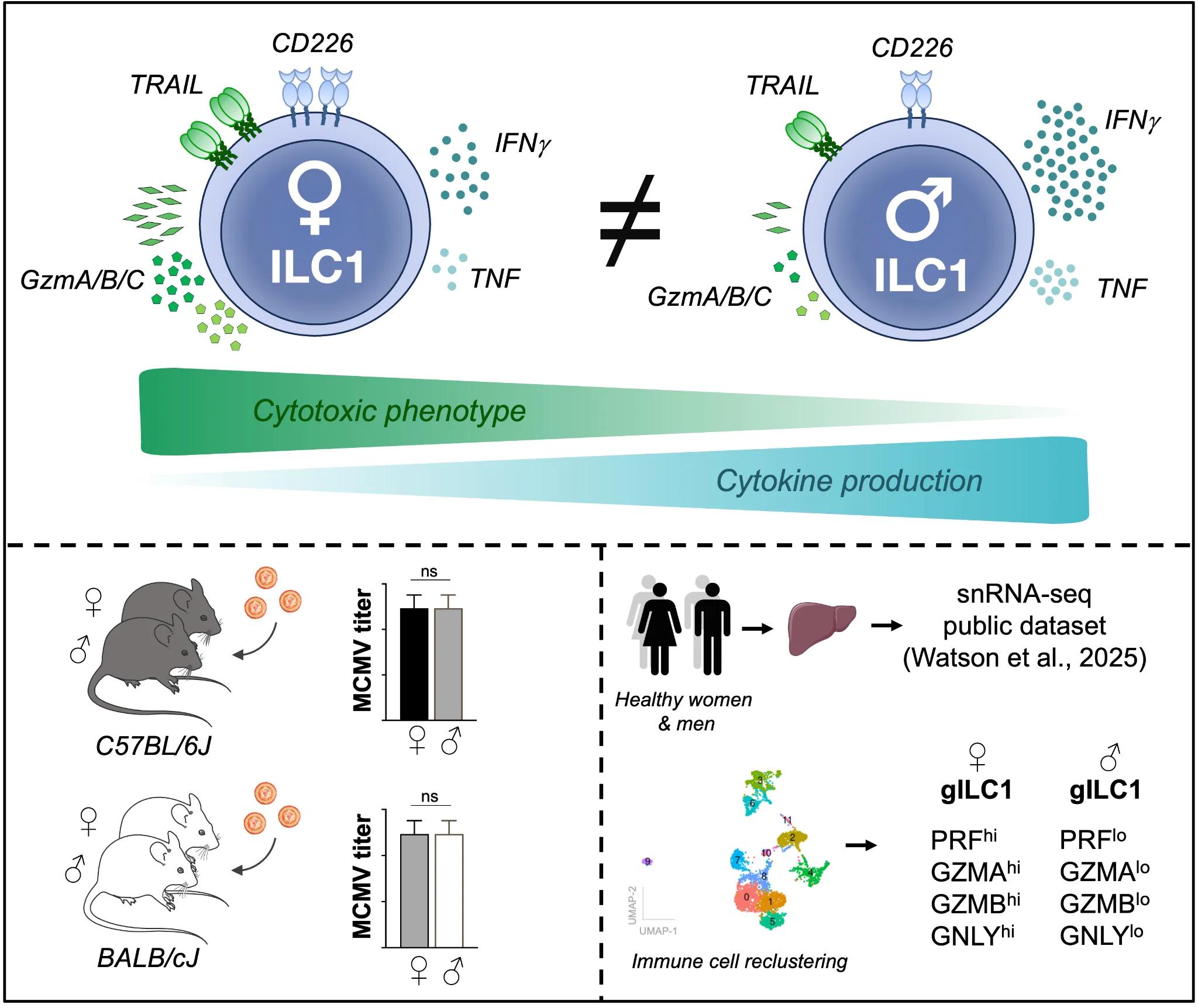
Abstract photo. Here, we investigated sex differences in tissue-resident type-I innate lymphoid cells (ILC1) in C57BL/6J and BALB/cJ mice. Adult females displayed stronger cytotoxic features, whereas males mounted a more inflammatory response during viral infection. Despite these differences, both sexes controlled MCMV infection equally well. Analysis of gILC1 transcriptome in healthy human liver confirmed the existence of sex-bias in liver-resident gILC1. This data suggests sex-biased ILC1 functions could influence immune-related pathologies.
Abstract
Sex-bias in the immune system is widely observed from insects to humans. While the evolutionary reasons for this are not clearly understood, the consequences for human health are more concrete. Women are generally more susceptible to autoimmune diseases, while men are more prone to develop cancer and infectious diseases. Furthering our understanding of the intrinsic biological differences in male and female immune systems should shed light upon sex-biased immune-related pathologies and potentially targeted therapeutics. In this context, we characterized the phenotype of group-I innate lymphoid cells (gILC1: conventional NK and ILC1) in male and female mice from C57Bl/6J and BALB/cJ strains. We showed that female ILC1 display higher basal activation levels with higher cytotoxic potential (TRAIL, GzmB, GzmC). This was only observed in adult mice, and not in young or older mice. Despite these differences, adult male mice displayed a similar ability to control mouse cytomegalovirus (MCMV) replication compared to female mice. This prompted us to investigate ILC1 response to infection. Male ILC1 still displayed lower granzyme expression compared to female ILC1 during MCMV infection, but produced greater amounts of inflammatory cytokines (IFNγ, TNF). Finally, we confirmed our findings in humans by analyzing a public dataset of single-nuclei RNA-seq, showing that liver-resident gILC1 express higher levels of cytotoxic molecules in healthy females compared to healthy males. Overall, we describe a sex-biased phenotype in ILC1 conserved in two strains of mice and mirrored in human liver gILC1, which may be relevant to human pathologies.
Keywords: cNK, human gILC1, ILC1, inflammation, MCMV, sex-bias
Introduction
It is well established that males and females display physiologically relevant, sex-dependent differences (Beery and Zucker, 2011, Lee, 2018, Woitowich et al., 2020, Zucker and Prendergrast, 2020, Plevkova et al., 2021). The immunology field is no exception, as females generally display stronger immune responses than males (Whitacre, 2001, Ngo et al., 2014, Klein and Flanagan, 2016, Qi et al., 2021, Dhakal et al., 2022). These differences between males and females are believed to be at least partially responsible for the stronger incidence of autoimmune diseases in females, and conversely, stronger incidence of cancer and susceptibility to infection in males. As such, having a deeper understanding of the differences between male and female immune systems is key to understanding the pathophysiology of a variety of diseases, and could lead to more fine-tuned therapies (Zucker and Prendergrast, 2020, Pala et al., 2022). In this context, we sought to investigate the implication of sex-bias on the phenotype and function of tissue-resident type-I innate lymphoid cells (ILC1) in two different strains of mice (B6J and BALB/cJ). Like conventional natural killer cells (cNK), ILC1 belong to the group-I innate lymphoid cells family (gILC1), characterized by the lack of genetically rearranged antigen receptors, expression of NK1.1 and/or NKp46, T-bet, a robust production of IFN𝛾, and cytotoxicity. We and others have shown that ILC1 are involved in anti-viral responses to MCMV (murine cytomegalovirus), anti-tumor responses, autoimmunity, and in non-alcoholic liver diseases (NALD) (Weizman et al., 2017, Picarda et al., 2019, Weizman et al., 2019, Cheng et al., 2023a, Evers et al., 2023, Hildreth et al., 2023, Zhang et al., 2023, Marrocco et al., 2025). While studies assessing the role of sex-bias for cNK have been published (Hou and Zheng, 1988, Zientarska et al., 2021, Cheng et al., 2023b), its role in regulating ILC1 remains unclear. ILC1 are resident mainly in non-lymphatic tissues and are well-studied in the liver (Fuchs, 2016, McFarland et al., 2021, Jaeger et al., 2024), but they are also present in the spleen and salivary glands (SG). Here, we compared male and female ILC1 phenotypes in both C57BL/6J (B6J) mice and BALB/cJ mice, focusing mainly on the liver, but also analyzing these other two tissues. We then assessed how sex-bias might impact the ability to control MCMV replication. Our results show that female ILC1 in the liver are more numerous and display higher activation levels (higher expression of CD226, TRAIL, GzmA, GzmB, GzmC) compared to male ILC1, suggesting higher cytotoxic potential. While there was very little sex-bias observed among gILC1 in the spleen, a profound change in the expression of TRAIL, GzmC, Eomes and Ly49H was observed in the salivary glands, all four proteins being drastically decreased in male gILC1 compared with female gILC1. However, these differences did not lead to superior control of MCMV replication in females. As we found that male ILC1 produce more cytokines during MCMV infection, we hypothesize this may counterbalance their lower apparent cytotoxic potential. Alternatively, other cell populations displaying unique phenotypes in males and females could shape ILC1-based differences in vivo. Finally, we analyzed a public dataset of snRNA-seq performed on healthy human livers from both sexes to see if these differences are conserved in humans.
Methods and Materials
Mice
Wild-type (WT) B6J CD45.2 (C57BL/6J), and BALB/cJ mice were purchased from Jackson Laboratories (Sacramento, CA, USA) and bred in-house. Unless stated otherwise, 8 to 12 week-old mice were used for all experiments. This study was carried out in strict accordance with the guidelines of Association for assessment and Accreditation of laboratory Animal Care (AAALAC) and National Institutes of Health (NIH). All animal protocols (AP00001034) used in this study, were approved by the Institutional Animal Care and Use Committee (IACUC) of the La Jolla Institute for immunology (LJI).
FACS analysis
Liver cells were harvested and processed as previously published (Picarda et al., 2019), and analyzed on an LSR-II Fortessa (BD) or a Cytek Aurora. Antibodies used were as follow: CD3 (17A2), CD19 (6D5), F4/80 (T45-2342), NK1.1 (PK136), NKp46 (29A1.4), TRAIL (N2B2), CD49a (Ha31/8), CD49b (DX5), CD200R (OX-110), CD226 (DX11), Ly49H (3D10), Eomes (Dan11mag), Prf (S16009A), GzmA (GzA-3G8.5), GzmB (QA16A02), GzmC (SFC1D8), IFNγ (XMG 1.2), TNF (MP6-XT22). Dead cells were stained with LD blue (Invitrogen). Fluorescently conjugated antibodies were purchased from BD Biosciences and BioLegend. Normalized MFI was used when pooling results from two independent experiments to avoid batch effects. The average MFI of the female group in each experiment was set as 100%, and individual values were expressed as relative percentage to this reference. For cytokine production analysis, total liver cells were incubated for 4h in presence of GolgiPlug (Brefeldin A), without any stimulus, and then stained for cytometry analysis.
MCMV infection
MCMV (Smith or K181 strains) was originally produced in 3T3 cells from cloned and sequenced BAC DNA (gift of B. Adler and Alec Redwood; Redwood et al., 2005) and was then amplified in vitro on 3T3 and titrated on MEF (murine embryonic fibroblast) cells (Verma et al., 2014). Male and female mice were infected i.p. with 106 pfu in 100µL of PBS and euthanized 4- or 12-days post infection. STOP mutations of the MCMV-Smith (or K181) BAC were performed in E. coli by en-passant (ET) mutagenesis, and successful mutagenesis was confirmed by PCR sequencing. The entire BAC was also sequenced to verify no additional mutations existed in the mutant BAC as compared to the WT BAC. Smithm166-STOP was previously used and described (Picarda et al., 2019). Smith∆m157 was a gift from Dr. Luka Cicin-Sain. All viruses induced a 0% mortality in both sexes in all the experiments performed.
MCMV quantification by qPCR
When harvested for qPCR, mouse tissues were cut in ~2-3mm3 pieces, placed in a screwed tube, immersed in liquid nitrogen, and subsequently stored at -80°C until further processing. DNA was purified using the NucleoSpin Tissue, Mini kit for DNA from cells and tissue (Macherey-Nagel, #740952.250), according to manufacturer instructions. Briefly, samples were allowed to thaw and mechanically dissociated using KIMBLE® Disposable PELLET PESTLE® Tissue Grinder (749540-0000) with SP Bel-Art ProCulture Cordless Homogenizer (#F65100-0000). Samples were then incubated and agitated overnight with Proteinase K at 56°C. Digested samples were then used for DNA extraction following kit instructions. Purified DNA was quantified using a NanoDrop, normalized at 50-100ng/µL, and stored at -20°C until further processing. The following primers and probes were used for qPCR analysis: gB_F: AAGGCGTGGACTAGCGATAA; gB_R: GCAGTCTAGTCGCTTTCTGC; gB_probe: AGCTAGACGACAGCCAACGCACCGA (FAM/ZEN™/3′ IB™FQ); Act_F: AGAGGGAAATCGTGCGTGAC; Act_R: AGGAAGAGGATGCGGCAGTG; Act_probe: CTGTGCTATGTTGCTCTAGACTTCGAG (FAM/ZEN™/3′ IB™FQ). gB primers were used and published previously (Sitnik et al., 2023). gB probe was adapted from another study (Vliegen et al., 2003). Both were purchased from IDT, and diluted in IDTE buffer, pH8. qPCR was performed with 2µL of sample DNA, 4.5µL of IDT PrimeTime Gene Expression MasterMix (2X) (#1055771), 300nM of each primer, 200nM of the probes, in a final volume of 9µL. Bio-rad CFX384 Real-time system, C1000 Touch Thermal Cycler were used. Cycles were as follow: 95°C 3min; [95°C x 5sec; 60°C x 30sec] for 45 cycles, according to IDT PrimeTime Master Mix instruction for fast cycling. A plasmid containing 1 copy of gB target sequence and 1 copy of Actine target sequence was serially diluted and used as standard for MCMV-genome copy quantification. The expected amplicon size (136bp for gB; 93bp for Actin) were confirmed by gel electrophoresis.
Human liver snRNA-seq analysis
A public dataset (Rahman et al., 2025, Watson et al., 2025) was retrieved from the GEO database in October 2025 (GSE210077). The 34 liver sample files (Liver-1 to Liver-35; no sample named Liver-12) were processed on Rstudio 4.2.2 via Seurat 4.2.2. Nuclei with low gene or UMI counts (<500) or elevated mitochondrial or ribosomal read content (>10%) were filtered out, consistent with the initial studies (Rahman et al., 2025, Watson et al., 2025). Filtered nuclei were merged, normalized, and scaled. The harmony package was used for batch correction across samples. Expression of XIST and Y-chromosomes genes (RPS4Y1, DDX3Y, KDM5D, EIF1AY, USP9Y, ZFY, UTY) were used to calculate an X-score and Y-score and assign a sex to each sample based on average nuclear expression. Total nuclei were clustered (resolution 0.1) and automatically cell-annotated using ScType (Ianevski et al., 2022, Rahman et al., 2025). “Hematopoietic cells” and “immune system cells” were extracted and re-clustered on their own (resolution 0.5). Samples 24–36 exhibited a pronounced batch effect relative to the first 22 samples, with >95% of nuclei segregating into distinct clusters despite batch correction, and were therefore excluded from all analyses. The following genes were used to calculate a gILC1-score: IFNG, TBX21, EOMES, ZNF683, NCR1, NCAM1, KLRB1, KLRC2, KLRF1, ITGA1, IL7R, CXCR6, PRF1, GZMB, GNLY, NKG7, KLRD1, FCGR3A.
Results
Female ILC1 display a stronger effector phenotype than male ILC1
To study sex-bias in ILC1, we analyzed ILC1 from the two most frequently used mouse strains in immunological research: B6J and BALB/cJ mice. Group-I ILCs (gILC1) display a rather conserved phenotype in both strains, despite two main differences: BALB/cJ gILC1 do not express NK1.1 or Ly49H and express a higher level of NKp46. In both strains, liver cNK and ILC1 can be discriminated based on the expression of Eomes and CD49b (cNK), and expression of CD49a and TRAIL (ILC1) (Picarda et al., 2019, Marrocco et al., 2025) (Figure S1A). We observed a consistent decrease of the absolute number of ILC1 in male B6J mice compared to female B6J mice (Figure S1B). In addition, female ILC1 displayed higher expression of CD226, GzmB, and GzmC compared to males ILC1 (Figure 1A). TRAIL, GzmA and Prf were also decreased in some experiments or showed similar expression levels, but were never increased in males (Figure 1A, Figure 2), suggesting that sex-bias in ILC1 impacts some effector molecules more than others. As biological observations in a given genetic background can differ, we sought to confirm this phenotypic sex-bias in ILC1 in a different mouse strain. B6J and BALB/cJ are known to have different sensitivity to infection and cancer due in part to different types of dominant immunity (type-1 immunity bias in B6J; type-2 immunity bias in BALB/cJ), which could impact the expression of gILC1 genes. Our analyses in BALB/cJ confirmed in a second genetic background that female ILC1 are more numerous (Figure S1C) and display higher expression levels of CD226, TRAIL, GzmA, GzmB, GzmC, and Prf (Figure 1B). Of note, cNK were also analyzed in parallel in every experiment. Sex-bias was also observed in cNK, albeit less consistently (e.g. increased GzmB, GzmC, and Prf in B6J females, but not in BALB/c females) (Figure S1D, Figure S1E). While female cNK do have statistically higher GzmA and Prf levels than male cNK, these minor differences may not have biological relevance (Figure S1E). Like ILC1, female cNK were more numerous in both mouse strains compared to male ILC1. This contradicts the usual view that males have more cNK (Cheng et al., 2023b), which might be due to different organs analyzed (spleen vs liver here). Collectively, these results suggest that female ILC1 (and to a lesser extent, cNK) might be better equipped to sense and activate in response to stress signals (CD226) and perform contact-dependent (TRAIL) or soluble (GzmA/B/C, Prf) cell cytotoxicity to control viral infection or tumor development.
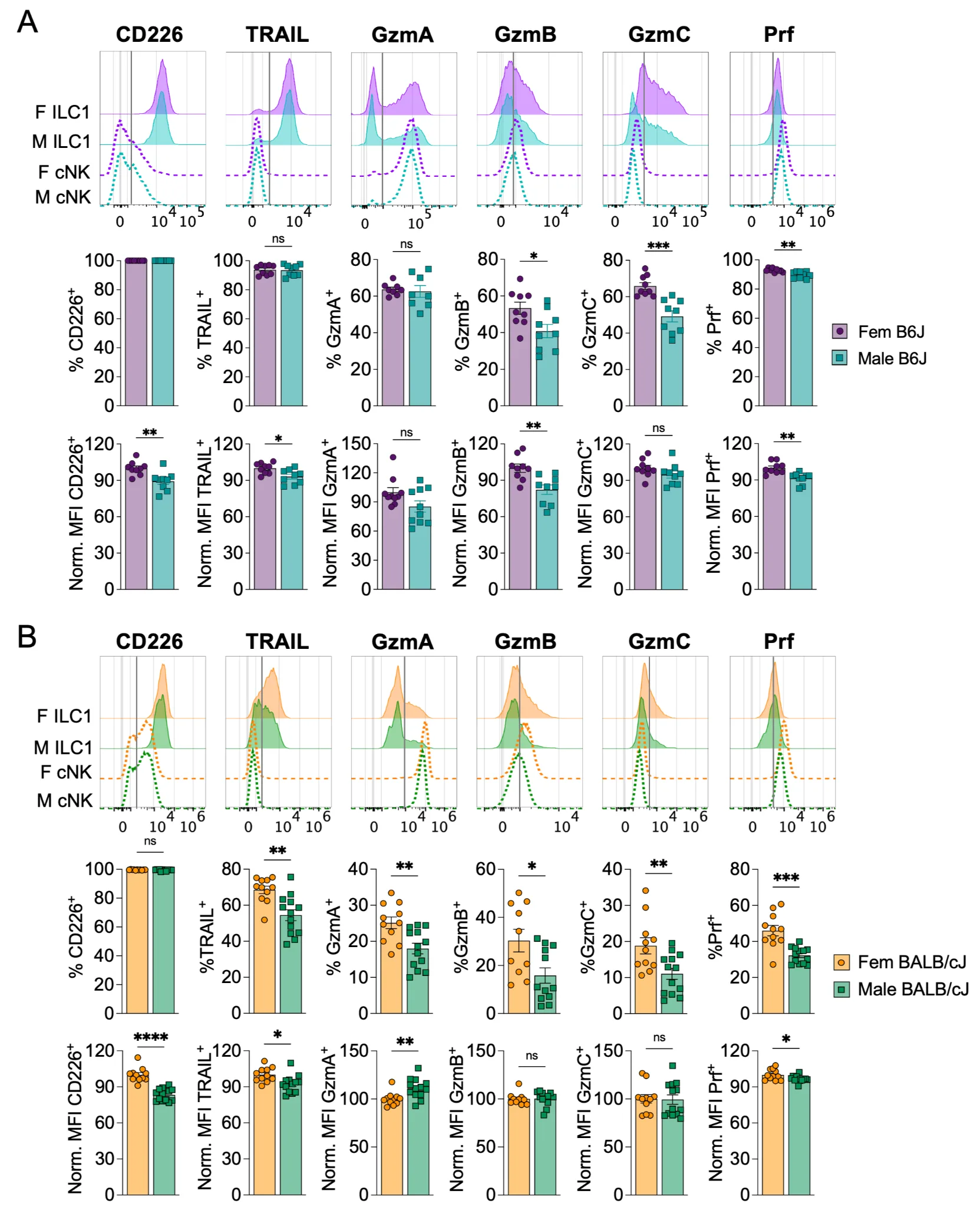
Figure 1. Female ILC1 display a more robust effector phenotype than male ILC1. Male and female adult mice were euthanized and perfused with PBS before harvesting the liver for cytometry analysis ex vivo. Representative stainings are shown for ILC1 (plain line) and cNK (dotted line) for all quantified markers. (A) Analysis performed on 10-weeks old C57BL/6J mice (B6J). Pool of 2 independent experiments (B) Analysis performed on 12-weeks old BALB/cJ mice. Pool of 2 independent experiments. Data were analyzed by unpaired t-test. Normality and homoscedasticity were assessed. Welch’s t-test was performed when samples showed heteroscedasticity. Mann-Whitney was performed when normality tests were negative. (ns: non-significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
Sex-bias in ILC1 peaks in adulthood
To further characterize potential sex-biases in ILC1, we asked whether the observed phenotypic differences existed at different ages. We assessed ILC1 phenotypes in B6J mice before sexual maturity (3 weeks), at early adulthood (12 weeks), and in older mice (14 months). As previously described (Asahi et al., 2023), ILC1 make up the majority of gILC1 in young animals, with their frequency and number decreasing in adulthood (Figure S2A). Notably, female mice contain more ILC1 than male mice during adulthood only, not in young or older mice (Figure S2A). TRAIL, GzmA, GzmB, and GzmC showed a strong age-dependent expression level in both sexes, overall decreasing with age. For example, TRAIL expression decreases between young and adult mice, and between adult and old mice, both in terms of percentage and MFI (Figure 2). We did not observe such striking and general age-dependent downregulation of these same markers in cNK. In fact, while GzmA MFI levels slightly decrease over time, perforin MFI levels and GzmB percentage increase with age in cNK (Figure S2B). Importantly, sex-biases in ILC1 were observed primarily in adult mice, with female ILC1 displaying higher levels of TRAIL and all three granzyme molecules assessed, confirming our previous observation. We also investigated gILC1 phenotype in the spleen and salivary glands of this cohort. Very few differences were found in the spleen for both cNK (Figure S2C, increased GzmB percentage in females) and ILC1 (Figure S3A, increased GzmC percentage in females). In the SG however, gILC1 were greatly impacted (Figure S3B). Since ILC1-vs-cNK phenotype varies dramatically by the tissue examined, especially in the SG, and no gating strategy allowed us to reproducibly discriminate them in all ages of mice, we analyzed total gILC1. Both ILC1-defining markers (TRAIL, GzmC) and cNK-defining markers (Eomes, Ly49H) were strongly impacted by both age and sex. These four markers are all identically expressed in young males and females, but are downregulated in adults and old males compared to females (Figure S3B). TRAIL and Ly49H expression is almost undetectable in adult males, while remaining highly expressed in females. Notably, Ly49H expression only drops in male gILC1 in the salivary glands, but not in the spleen or the liver. These results show that gILC1 phenotypes are sex, age, and tissue specific, and suggest that heightened activation in females or reduced activation in males could contribute to pathophysiologies in adult mice.
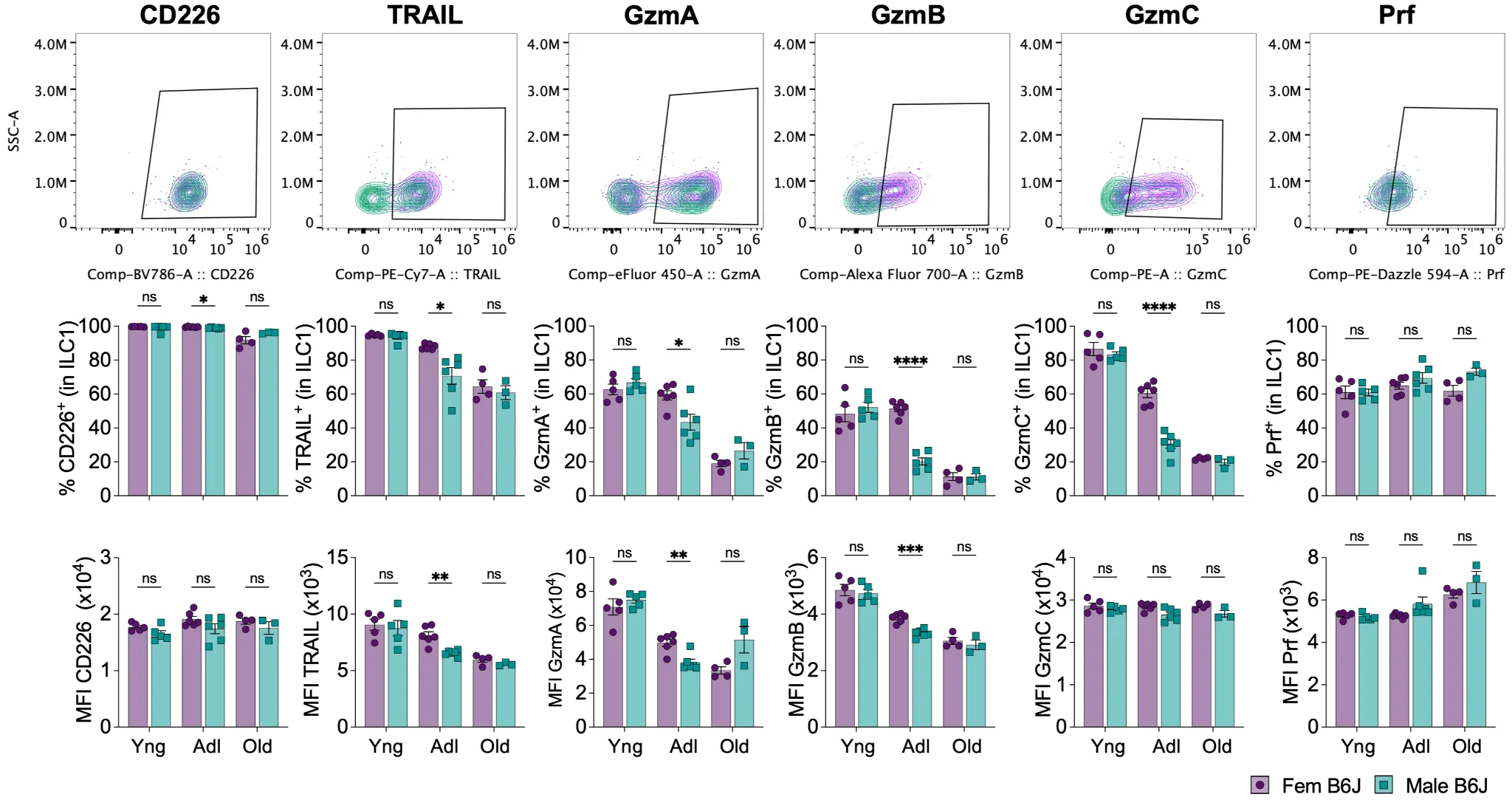
Figure 2. Sex-bias in ILC1 peaks in adulthood. Young (3w), adult (12w), and old (14m) male and female B6J mice were euthanized and perfused with PBS before harvesting the liver for cytometry analysis ex vivo. The upper panel shows representative plots and gates from adult females (purple) and adult males (green). Yng: young; Adl: adult. Data were analyzed by multiple unpaired t-tests, with Holm-Šídák correction. (ns: non-significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
Males and females show similar ability to control MCMV infection
Given the observed ILC1 sex-bias in adult mice, we next investigated if these differences would lead to difference in the ability to control MCMV infection, a virus known to trigger ILC1 activation and effector functions. We infected male and female B6J or BALB/cJ mice with various MCMV strains. First, we infected mice with MCMV-Smith and assessed the viral load in various tissues by qPCR at day 4 and day 12 post infection. Contrary to what we observed for phenotypic differences in both B6J and BALB/cJ mice, male and female mice showed similar viral load in the liver (Figure 3A), spleen, lungs, and salivary glands (Figure S4A, Figure S4B).
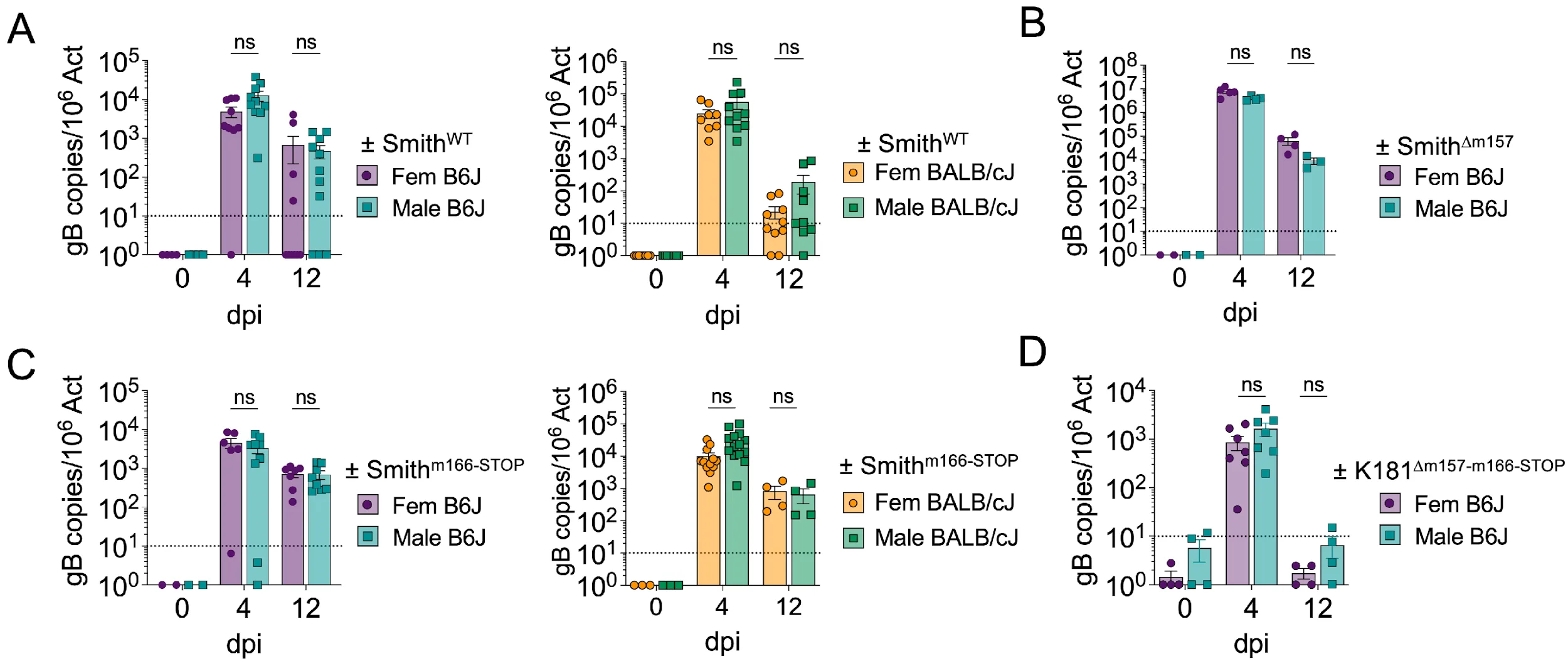
Figure 3. Males and females show similar ability to control MCMV infection. 8-to-12 week-old male and female adult mice were infected with various strains of virus and euthanized 4- and 12-days post infection (dpi). In each case, mice were infected through intraperitoneal injection of 106pfu of in vitro produced MCMV, diluted in 100µL of PBS. (A) Infection of B6J and BALB/cJ mice with untouched Smith-MCMV. Both graphs are a pool of 2 experiments. (B) Infection of B6J mice with m157-deficient Smith-MCMV. (C) Infection of B6J and BALB/cJ mice with m166-deficient Smith-MCMV. Both graphs are a pool of 2 experiments. (D) Infection of B6J mice with m157-deficient m166-deficient K181-MCMV. Data were analyzed by multiple unpaired t-tests, with Holm-Šídák correction. (ns: non-significant).
In B6J mice, cNK cells express Ly49H which binds to the MCMV protein m157 and leads to strong expansion and activation of cNK cells. We thought this potent cNK cell response could hinder an ILC1-dependent sex-bias effect during MCMV infection, and thus infected B6J mice with a mutant virus lacking m157 (Smith∆m157). As expected, the viral load in the liver (Figure 3B), spleen, lungs, and salivary glands (Figure S4C) were increased after infection with Smith∆m157 compared to SmithWT. However, males and females still showed similar ability to control Smith∆m157 infection. We previously demonstrated that MCMV can inhibit ILC1-mediated TRAIL immune control by downregulating the TRAIL-DR (Death Receptor) on the surface of infected cells via the viral protein m166 (Verma et al., 2014, Picarda et al., 2019). As this would potentially impair ILC1 responses and could hide the effect of ILC1-dependent sex-bias, we infected B6J and BALB/cJ mice with a mutant virus lacking m166: Smithm166-STOP. However, males and females still showed equivalent ability to control the infection (Figure 3C). Finally, we performed a similar experiment using a K181 virus lacking both m166 and m157 proteins. As before, we observed no difference in MCMV∆m157m166stop replication between males and females. In summary, the phenotypic differences we observe between male and female ILC1 do not lead to differences in the ability of the mice to restrict MCMV replication, even in the absence of robust m157-dependent cNK responses and/or inhibition of TRAIL-dependent killing.
Male ILC1 produce higher cytokines levels than female ILC1
While male ILC1 showed a weaker effector phenotype at steady-state compared to female ILC1, perhaps they respond more robustly in response to MCMV infection, accounting for similar MCMV control. To test this, we infected male and female B6J mice with MCMV-SmithWT and analyzed ILC1 from the liver at day 4 post infection. Notably, female ILC1 were still more numerous than male ILC1 (Figure 4A). In addition, CD226, TRAIL, GzmB, and GzmC were all still more highly expressed by female ILC1 compared to male ILC1 (Figure 4A). However, after a 4h incubation with brefeldin A, we measured a higher IFNγ and TNF production by male ILC1 compared to their female counterparts (Figure 4B), suggesting that ILC1 sex-bias is situation-dependent. The cytokine production by cNK did not differ between males and females (Figure S5). Importantly, the same phenotypic differences were also observed at earlier time-point (36h post infection, data not shown). However, while similar trends were observed for cytokine production, no significant differences were observed at that time-point (data not shown).
Sex bias in liver gILC1 in human
To investigate the translational aspect of our findings, we analyzed a publicly available dataset of single-nuclei RNA-seq performed on liver samples from 34 healthy human donors (GSE210077). To assign a sex to each sample, we calculated a XIST-score and Y-score (Figure S6A), revealing 21 female samples and 14 males samples. Samples 9 and 17 were excluded due to low nuclei count after QC (Figure S6B).
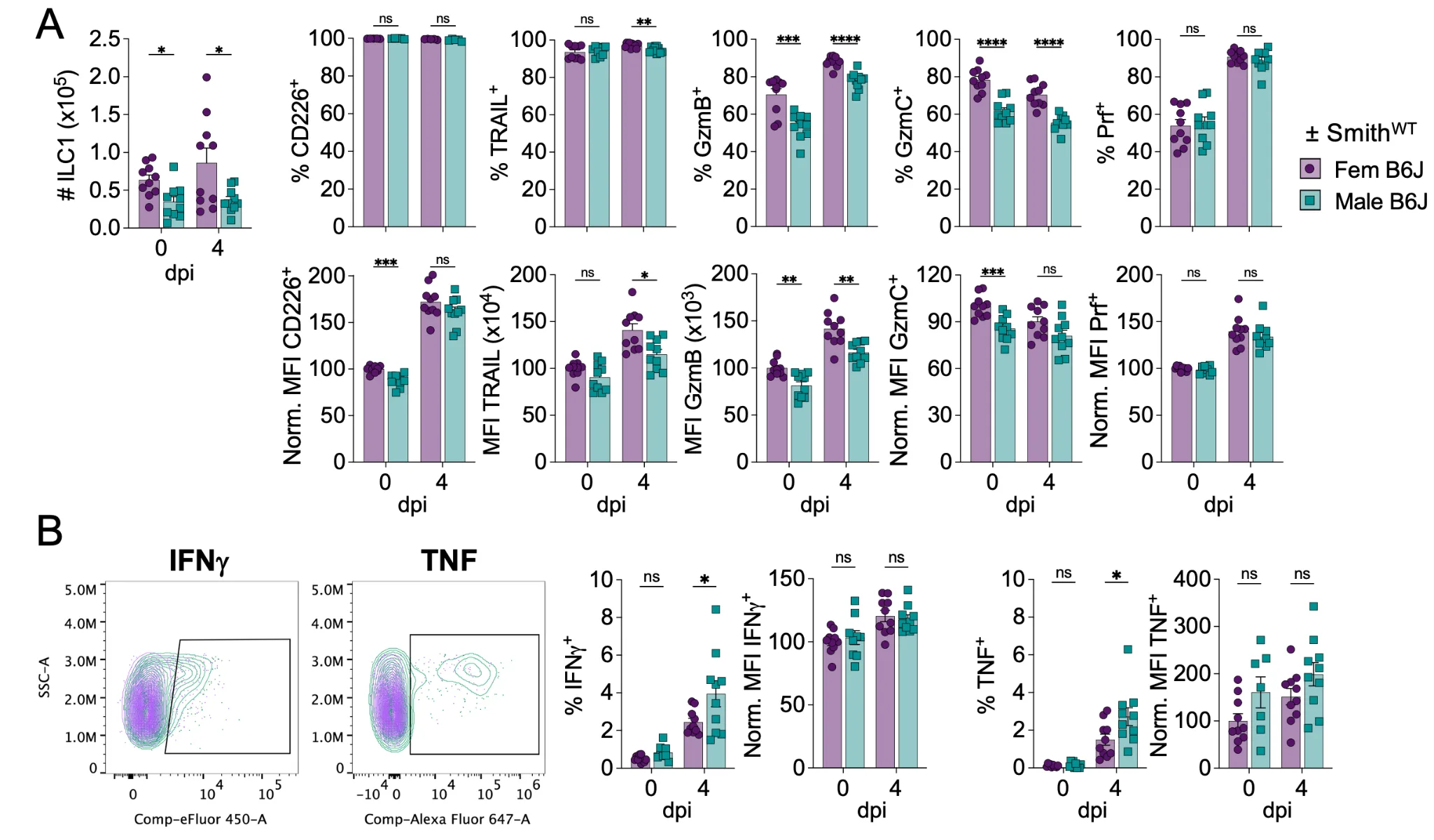
Figure 4. Male ILC1 produce more cytokines than female ILC1. Male and female B6J 8-weeks old mice were infected with Smith-MCMV (i.p., 106pfu) and euthanized 4-days post infection (dpi), together with non-infected control mice. Mice were perfused with PBS before harvesting the liver for cytometry analysis ex vivo (A) and following in vitro incubation with GolgiPlug (B). Data were analyzed by multiple unpaired t-tests, with Holm-Šídák correction. (ns: non-significant, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001).
Despite running harmony for batch-correction, samples 24 to 36 showed a critical batch-effect compared to the first 22 samples and were thus excluded. After QC and sample exclusion, we were left with 114,243 nuclei to analyze, from 11 female donors and 8 male donors. Automated annotation of those nuclei through ScType identified 7 populations (Figure 5A), of which “Hematopoietic cells” and “immune system cells” were selected and re-clustered on their own (Figure 5B). This led to the identification of 12 immune cell clusters. Looking at genes specific for T cells (CD3D, TRAC, THEMIS), gILC1 (NCR1, NKG7, KLRD1), B cells (CD79A, CD79B), monocytes (LYZ, S100A8), macrophages (MARCO, C1QA), dendritic cells (ITGAX, CLEC9A), or neutrophils (CSF3R), cluster 2 and 7 seemed to belong to the gILC1 family (Figure S6C).
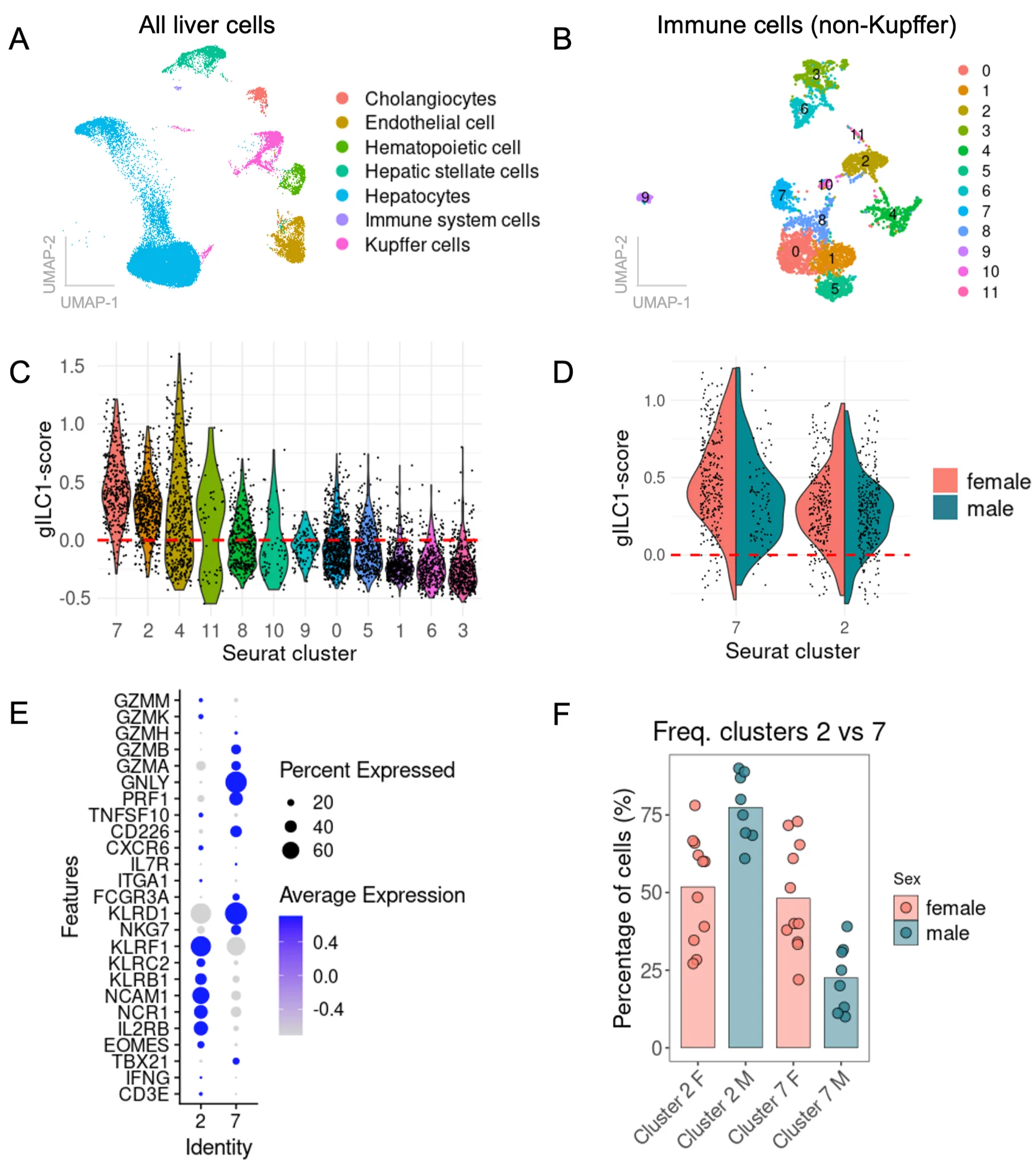
Figure 5. Sex-bias among human gILC1 in the liver. Analysis of a published snRNA-seq dataset on healthy human liver (GSE210077). (A) UMAP and clustering of all liver cells annotated with ScType. (B) UMAP and clustering of immune cells only, without Kupffer cells. (C) gILC1-signature score among immune clusters, based on the expression of 18 genes. (D) gILC1-signature score among clusters 2 and 7, stratified by sex. E expression of key gILC1 genes within cluster 2 compared to cluster 7. F Relative frequency of cluster 2 and 7 within females and males.
To validate this, we calculated an gILC1-signature score based on the expression of 18 genes related to gILC1 (Figure S6D, Figure 5C). Cluster 7 and 2 are the only clusters with >90% of the nuclei showing a positive score (Figure 5C). While cluster 4 also has a relatively high score for gILC1-signature genes, this cluster shows expression of CD3E, CD3D, TRAC and THEMIS, and thus likely contains NKT cells (Figure S6C). We focused on cluster 2 and 7 and re-analyzed the gILC1-signature score, this time comparing by sex (Figure 5D). No drastic differences were observed between males and females, even though cluster 7 in males tends to have a slightly lower score than cluster 7 in females. In terms of phenotype, cluster 2 shows more of an cNK-like signature, expressing higher levels of EOMES, NCAM1 (CD56), KLRB1, KLRF1 (NKp80), but express low level of cytotoxic molecules (Figure 5E). Cluster 7 shows no EOMES expression, but expresses TBX21, CD226, NKG7, KLRD1 (CD94), and a high level of cytotoxic molecules PRF1, GNLY, GZMA and GZMB. No cluster matched the LIN-IL7R+TBX21+ signature described in the literature for human ILC1. Importantly, when looking at count (Figure S6E) and relative proportion of cluster 2 and 7 among total gILC1 in males and females (Figure 5F), we observed a strong bias toward the non-cytotoxic cluster 2 in males, containing way less cells in the cytotoxic cluster 7. These results suggest that like in mice, human gILC1 might have stronger cytotoxic potential in females compared to males.
Discussion
Intrinsic differences between women and men’s biological processes have been acknowledged, analyzed, and characterized in many scientific research fields. While immunological research is now closing the gap, it has historically been a sex-agnostic biological field (Beery and Zucker, 2011, Klein et al., 2016, Woitowich et al., 2020). Given the wide array of immune related pathologies showing a bias toward males or females, it is crucial that we deepen our understanding of these sex-specific differences. Here we analyzed the phenotype of ILC1 and cNK in male and female mice from 2 different genetic backgrounds (C57BL/6J, and BALB/cJ). We showed that female liver ILC1 displayed a stronger effector phenotype in both strains of mice, and that among B6J mice this was particularly profound in adult mice compared to very young or >1 year old animals. Importantly, sex-bias among gILC1 varied greatly across tissues, with little variation between males and females in the spleen, but large differences within the salivary glands (Figure S2, Figure S3). This, together with the fact that gILC1 phenotypes evolve with age (Figure 2, Figure S2, Figure S3), suggest that gILC1 are highly susceptible to environmental cues. Multiple studies have highlighted that sex-bias is influenced by genetic differences (genes present on Y chromosome), epigenetic (second X chromosome inactivation, or lack thereof), microbiota, or hormonal levels (Yurkovetskiy et al., 2013, Ngo et al., 2014, Blanquart et al., 2021, Heydari et al., 2022, Cheng et al., 2023b, Dou et al., 2024). It is likely that several of these factors contribute to the phenotypes described here. Due to the specific age-window where ILC1 sex-bias was more potent in our study and in accordance with literature regarding sex-bias in ILC2 (Laffont et al., 2017), the observed differences between male and female ILC1 may be secondary to hormonal differences. Further investigations following mice gonadectomy would be informative. Importantly, while consistent, the sex-biased phenotype in ILC1 was not replicated in every experiment. We currently do not know what causes this variability, but (i) It cannot be explained by differences in the age of the adult mice. (ii) Immune variation recently described based on circadian-rhythms cannot explain it either, as mice necropsy were always performed at the same time. (iii) As males tend to fight, we think intra-experimental variability could be partially due to variation in stress level in the different batches of mice. (iv) Additionally, if these differences are indeed dependent on sexual hormone levels, variability in the female hormonal cycle timing could be expected to contribute. Of note, future studies on sex-bias and ILC1 might be better suited within BALB/cJ mice, as the differences between males and females seem more pronounced and consistent in this mouse strain (Figure 1).
Another important discrepancy is the weak sex-bias among cNK seen in our studies, which only partially reconciles with previously published work (Cheng et al., 2023b). cNK are generally detected in higher numbers in males compared to females in both the spleen of mice (Cheng et al., 2023b) or in human PBMC (Huang et al., 2021). We could not confirm those findings in our experiments when looking at the liver (Figure S1) or the spleen (Figure S2). Such differences could be due to tissue processing methods, organ-dependent differences, and the low number of animals analyzed in the spleen in our study. Importantly, the liver and splenic cNK population studied here excludes ILC1 cells, while the vast majority of published studies in splenocytes study the bulk CD3+NK1.1+ population, containing both cNK and ILC1. Finally, we show that males cNK produce equivalent levels of cytokines in response to MCMV, while other studies showed increased cytokine production in female cNK (Cheng et al., 2023b). These seemingly discordant results could emanate from our use of MCMV as an activating agent in vivo, while others used total gILC1 stimulated in vitro with IL-12/IL-15 (Cheng et al., 2023b). Ultimately, however, both our and other studies consistently show higher GzmB expression in female gILC1.
Despite higher activation levels and cytotoxic potential of female ILC1, males showed similar ability to control MCMV replication. This was true in two mouse genetic backgrounds tested, using a variety of MCMV strains and mutants. This might be due to male ILC1 producing more inflammatory cytokines compared to females, counterbalancing direct antiviral effector functions, or perhaps other cells are involved in vivo. Given the observed sex-bias in cytotoxic phenotype, assessing the killing ability of male vs female ILC1 to confirm whether the observed phenotypic difference does translate to functional differences would provide important information. Critically, the use of a public dataset of snRNA-seq allowed us to extrapolate our findings to healthy human liver gILC1. While we did not observe a population displaying the reported ILC1-phenotype (Lin-IL7R+T-BET+), we saw 2 populations with high expression of gILC1 signature genes. Cluster 2 – overrepresented in males – expresses EOMES and low levels of cytotoxic molecules, while cluster 7 – overrepresented in females – expresses TBX21 and high levels of cytotoxic molecules. This suggests that our findings in mice translate to people and could be relevant to human health. Further study of inflammatory conditions such as NASH (Nonalcoholic Steatohepatitis), where ILC1-derived IFN-γ is a critical component (13), and ILC1-mediated killing of tumor cells (Kansler et al., 2022), will provide important insight into how sex-biases in ILC1 may regulate different outcomes in men and women.
Acknowledgments
We would like to thank Catia Simoes, Juan-Pablo Romero, and Suzie Alarcón for their help in analyzing the human transcriptomic dataset, together with La Jolla institute IT infrastructure providing access to the LJI cluster.
Author Contributions
Remi Marrocco: Conceptualization, Data Curation, Formal Analysis, Investigation, Methodology, Supervision, Visualization, Writing – Original Draft, Writing – Review & Editing
Meghan Nguyen: Investigation
Andrea Balcan: Investigation, Visualization, Writing – Original Draft
Kaia Robinson: Investigation
Chris A. Benedict: Conceptualization, Funding acquisition, Methodology, Supervision, Writing – Original Draft, Writing – Review & Editing
Data Availability
The data and materials are available on request.
Supplementary Information
Supplementary information can be found here.
Transparent Peer Review
Results from the Transparent Peer Review can be found here.
Recommended Citation
Marrocco, R., M. Nguyen, A. Balcan, K. Robinson, and C. A. Benedict. 2026. Sex biases in ILC1 phenotypes do not alter immune control of cytomegalovirus. Stacks Journal: 26004. https://doi.org/10.60102/stacks-26004
References
Asahi, T., S. Abe, G. Cui, A. Shimba, T. Nabekura, H. Miyachi, S. Kitano, K. Ohira, J. M. Dijkstra, M. Miyazaki, A. Shibuya, H. Ohno, and K. Ikuta. 2023. Liver type 1 innate lymphoid cells lacking IL-7 receptor are a native killer cell subset fostered by parenchymal niches. eLife 12. https://doi.org/10.7554/eLife.84209.
Beery, A. K., and I. Zucker. 2011. Sex bias in neuroscience and biomedical research. Neuroscience & Biobehavioral Reviews 35: 565–572. https://doi.org/10.1016/j.neubiorev.2010.07.002.
Blanquart, E., S. Laffont, and J.-C. Guery. 2021. Sex hormone regulation of innate lymphoid cells. Biomedical Journal 44: 144–156. https://doi.org/10.1016/j.bj.2020.11.007.
Cheng, C., Q. Zhang, Y. Li, J. Jiang, L. Xie, H. Shen, D. Wu, H. Zhang, H. Zhang, X. Wang, H. Wu, J. Xu, L. Gui, B. Li, C. Ju, H. Peng, S. Yin, and L. Xu. 2023a. Interplay between liver type 1 innate lymphoid cells and NK cells drives the development of alcoholic steatohepatitis. Cellular and Molecular Gastroenterology and Hepatology 15: 261–274. https://doi.org/10.1016/j.jcmgh.2022.09.010.
Cheng, M. I., J. H. Li, L. Riggan, B. Chen, R. Y. Tafti, S. Chin, F. Ma, M. Pellegrini, H. Hrncir, A. P. Arnold, T. E. O’Sullivan, and M. A. Su. 2023b. The X-linked epigenetic regulator UTX controls NK cell-intrinsic sex differences. Nature Immunology 24: 780–791. https://doi.org/10.1038/s41590-023-01463-8.
Dhakal, S., S. Chaulagain, and S. L. Klein. 2022. Sex biases in infectious diseases research. Journal of Experimental Medicine 219: e20211486. https://doi.org/10.1084/jem.20211486.
Dou, D. R., Y. Zhao, J. A. Belk, Y. Zhao, K. M. Casey, D. C. Chen, R. Li, B. Yu, S. Srinivasan, B. T. Abe, K. Kraft, C. Hellström, R. Sjöberg, S. Chang, A. Feng, D. W. Goldman, A. A. Shah, M. Petri, L. S. Chung, D. F. Fiorentino, E. K. Lundberg, A. Wutz, P. J. Utz, and H. Y. Chang. 2024. Xist ribonucleoproteins promote female sex-biased autoimmunity. Cell 187: 733–749. https://doi.org/10.1016/j.cell.2023.12.037.
Evers, B. D. G., M. Hils, C. Heuser, I. M. Hölge, D. Argiriu, Y. Skabytska, S. Kaesler, C. Posch, P. A. Knolle, and T. Biedermann. 2023. Inflammatory cues direct skin-resident type 1 innate lymphoid cells to adopt a psoriasis-promoting identity. JID Innovations 3: 100204. https://doi.org/10.1016/j.xjidi.2023.100204.
Fuchs, A. 2016. ILC1s in tissue inflammation and infection. Frontiers in Immunology 7: 104. https://doi.org/10.3389/fimmu.2016.00104.
Heydari, R., Z. Jangravi, S. Maleknia, M. Seresht-Ahmadi, Z. Bahari, G. H. Salekdeh, and A. Meyfour. 2022. Y chromosome is moving out of sex determination shadow. Cell Bioscience 12: 4. https://doi.org/10.1186/s13578-021-00741-y.
Hildreth, A. D., E. T. Padilla, R. Y. Tafti, A. R. Legala, and T. E. O’Sullivan. 2023. Sterile liver injury induces a protective tissue-resident cDC1–ILC1 circuit through cDC1-intrinsic cGAS–STING-dependent IL-12 production. Cell Reports 42: 112141. https://doi.org/10.1016/j.celrep.2023.112141.
Hou, J., and W. F. Zheng. 1988. Effect of sex hormones on NK and ADCC activity of mice. International Journal of Immunopharmacology, 10: 15–22.
Huang, Z., B. Chen, X. Liu, H. Li, L. Xie, Y. Gao, R. Duan, Z. Li, J. Zhang, Y. Zheng, and W. Su. 2021. Effects of sex and aging on the immune cell landscape as assessed by single-cell transcriptomic analysis. Proceedings of the National Academy of Sciences USA 118: e2023216118. https://doi.org/10.1073/pnas.2023216118.
Ianevski, A., A. K. Giri, and T. Aittokallio. 2022. Fully-automated and ultra-fast cell-type identification using specific marker combinations from single-cell transcriptomic data. Nature Communications 13: 1246. https://doi.org/10.1038/s41467-022-28803-w.
Jaeger, N., A. U. Antonova, D. Kreisel, F. Roan, E. Lantelme, S. F. Ziegler, M. Cella, and M. Colonna. 2024. Diversity of group 1 innate lymphoid cells in human tissues. Nature Immunology 25: 1460–1473. https://doi.org/10.1038/s41590-024-01885-y.
Kansler, E. R., S. Dadi, C. Krishna, B. G. Nixon, E. G. Stamatiades, M. Liu, F. Kuo, J. Zhang, X. Zhang, K. Capistrano, K. A. Blum, K. Weiss, R. M. Kedl, G. Cui, K. Ikuta, T. A. Chan, C. S. Leslie, A. A. Hakimi, and M. O. Li. 2022. Cytotoxic innate lymphoid cells sense cancer cell-expressed interleukin-15 to suppress human and murine malignancies. Nature Immunology 23: 904–915. https://doi.org/10.1038/s41590-022-01213-2.
Klein, S. L., and K. L. Flanagan. 2016. Sex differences in immune responses. Nature Reviews Immunology 16: 626–638. https://doi.org/10.1038/nri.2016.90.
Laffont, S., E. Blanquart, M. Savignac, C. Cénac, G. Laverny, D. Metzger, J. P. Girard, G. T. Belz, L. Pelletier, C. Seillet, and J.-C. Guéry. 2017. Androgen signaling negatively controls group 2 innate lymphoid cells. Journal of Experimental Medicine 214: 1581–1592. https://doi.org/10.1084/jem.20161807.
Lee, S. K. 2018. Sex as an important biological variable in biomedical research. BMB Reports 51: 167–173. https://doi.org/10.5483/BMBRep.2018.51.4.034.
Remi M., E. Lucero-Meza, and C. A. Benedict. 2025. Type I interferon regulation of group I ILC subsets during both homeostasis and cytomegalovirus infection. The Journal of Immunology 214:1733–1740. https://doi.org/10.1093/jimmun/vkaf051.
McFarland, A. P., A. Yalin, S. Y. Wang, V. S. Cortez, T. Landsberger, R. Sudan, V. Peng, H. L. Miller, B. Ricci, E. David, R. Faccio, I. Amit, and M. Colonna. 2021. Multi-tissue single-cell analysis deconstructs the complex programs of mouse natural killer and type 1 innate lymphoid cells in tissues and circulation. Immunity 54: 1320–1337. https://doi.org/10.1016/j.immuni.2021.03.024.
Ngo, S. T., F. J. Steyn, and P. A. McCombe. 2014. Gender differences in autoimmune disease. Frontiers in Neuroendocrinology 35: 347–369. https://doi.org/10.1016/j.yfrne.2014.04.004.
Pala, L., T. De Pas, C. Catania, G. Giaccone, A. Mantovani, S. Minucci, G. Viale, R. D. Gelber, and F. Conforti. 2022. Sex and cancer immunotherapy: Current understanding and challenges. Cancer Cell 40: 695–700. https://doi.org/10.1016/j.ccell.2022.06.005.
Picarda, G., R. Ghosh, B. McDonald, S. Verma, N. Thiault, R. E. Morabiti, T. S. Griffith, and C. A. Benedict. 2019. Cytomegalovirus evades TRAIL-mediated innate lymphoid cell 1 defenses. Journal of Virology 93. https://doi.org/10.1128/jvi.00617-19.
Plevkova, J., M. Brozmanova, J. Harsanyiova, M. Sterusky, J. Honetschlager, and T. Buday. 2021. Various aspects of sex and gender bias in biomedical research. Physiological Research 69: S367–S378. https://doi.org/10.33549/physiolres.934593.
Qi, S., C. Ngwa, D. A. Morales Scheihing, A. Al Mamun, H. W. Ahnstedt, C. E. Finger, G. D. Colpo, R. Sharmeen, Y. Kim, H. A. Choi, L. D. McCullough, and F. Liu. 2021. Sex differences in the immune response to acute COVID-19 respiratory tract infection. Biology of Sex Differences 12: 66. https://doi.org/10.1186/s13293-021-00410-2.
Rahman, R. U., E. T. Epstein, S. Murphy, L. Amir-Zilberstein, C. McCabe, T. Delorey, H. Koene, L. Fernandes, K. K. Tanabe, M. Qadan, C. Ferrone, D. L. Berger, A. Shih, J. Deguine, and A. C. Mullen. 2025. Single-cell transcriptomics reveals the impact of sex and age in the healthy human liver. JHEP Reports: 101773. http://dx.doi.org/10.1016/j.jhepr.2026.101773.
Redwood, A. J., M. Messerle, N. L. Harvey, C. M. Hardy, U. H. Koszinowski, M. A. Lawson, and G. R. Shellam. 2005. Use of a murine cytomegalovirus K181-derived bacterial artificial chromosome as a vaccine vector for immunocontraception. Journal of Virology 79: 2998–3008. https://doi.org/10.1128/JVI.79.5.2998-3008.2005.
Sitnik, K. M., F. Krstanović, N. Gödecke, U. Rand, T. Kubsch, H. Maaß, Y. Kim, I. Brizić, and L. Čičin-Šain. 2023. Fibroblasts are a site of murine cytomegalovirus lytic replication and Stat1-dependent latent persistence in vivo. Nature Communications 14: 3087. https://doi.org/10.1038/s41467-023-38449-x.
Verma, S., A. Loewendorf, Q. Wang, B. McDonald, A. Redwood, and C. A. Benedict. 2014. Inhibition of the TRAIL death receptor by CMV reveals its importance in NK cell-mediated antiviral defense. PLoS Pathogens 10: e1004268. https://doi.org/10.1371/journal.ppat.1004268.
Vliegen, I., S. Herngreen, G. Grauls, C. Bruggeman, and F. Stassen. 2003. Improved detection and quantification of mouse cytomegalovirus by real-time PCR. Virus Research 98: 17–25. https://doi.org/10.1016/j.virusres.2003.08.009.
Watson, B. R., B. Paul, R. U. Rahman, L. Amir-Zilberstein, Å. Segerstolpe, E. T. Epstein, S. Murphy, L. Geistlinger, T. Lee, A. Shih, J. Deguine, R. J. Xavier, J. R. Moffitt, and A. C. Mullen. 2025. Spatial transcriptomics of healthy and fibrotic human liver at single-cell resolution. Nature Communications 16: 319. https://doi.org/10.1038/s41467-024-55325-4.
Weizman, O. E., N. M. Adams, I. S. Schuster, C. Krishna, Y. Pritykin, C. Lau, M. A. Degli-Esposti, C. S. Leslie, J. C. Sun, and T. E. O’Sullivan. 2017. ILC1 confer early host protection at initial sites of viral infection. Cell 171: 795–808. https://doi.org/10.1016/j.cell.2017.09.052.
Weizman, O. E., E. Song, N. M. Adams, A. D. Hildreth, L. Riggan, C. Krishna, O. A. Aguilar, C. S. Leslie, J. R. Carlyle, J. C. Sun, and T. E. O’Sullivan. 2019. Mouse cytomegalovirus-experienced ILC1s acquire a memory response dependent on the viral glycoprotein m12. Nature Immunology 20: 1004–1011. https://doi.org/10.1038/s41590-019-0430-1.
Whitacre, C. C. 2001. Sex differences in autoimmune disease. Nature Immunology 2: 777–780. https://doi.org/10.1038/ni0901-777.
Woitowich, N. C., A. Beery, and T. Woodruff. 2020. A 10-year follow-up study of sex inclusion in the biological sciences. eLife 9. https://doi.org/10.7554/eLife.56344.
Yurkovetskiy, L., M. Burrows, A. A. Khan, L. Graham, P. Volchkov, L. Becker, D. Antonopoulos, Y. Umesaki, and A. V. Chervonsky. 2013. Gender bias in autoimmunity is influenced by microbiota. Immunity 39: 400–412. https://doi.org/10.1016/j.immuni.2013.08.013.
Zhang, Y., S. Ma, T. Li, Y. Tian, H. Zhou, H. Wang, and L. Huang. 2023. ILC1-derived IFN-γ regulates macrophage activation in colon cancer. Biology Direct 18: 56. https://doi.org/10.1186/s13062-023-00401-w.
Zientarska, A., J. Mikuła-Pietrasik, M. Kaczmarek, A. Witkowska, B. Rozpłochowski, A. Kowala-Piaskowska, K. Książek, J. Żeromski, and I. Mozer-Lisewska. 2021. Gender as a factor affecting NK cell activity in patients successfully treated for chronic hepatitis C with direct-acting antivirals. Central European Journal of Immunology 46: 481–491. https://doi.org/10.5114/ceji.2021.109846.
Zucker, I., and B. J. Prendergast. 2020. Sex differences in pharmacokinetics predict adverse drug reactions in women. Biology of Sex Differences 11: 32. https://doi.org/10.1186/s13293-020-00308-5.
Open Access
Peer-Reviewed
Creative Commons
Accepted: 09 February 2026
Published: 27 March 2026
Funding for the study was provided under grant number AI101423 “The role of TRAIL in immune control of cytomegalovirus.”
Conflicts of Interest:
The authors have no conflicts of interest with the research and results in this manuscript.







